Speak the same language
Last updated on 2026-04-14 | Edit this page
Overview
Questions
- What are data descriptions?
- How can data descriptions be reused?
- Are there standard ways to create them?
- What is the relationship between data descriptions and linked data?
Objectives
- Recognize that different communities use different names for data descriptions.
- Learn how to build machine-friendly data description files.
- Understand why reusing ontology terms improves interoperability.
FAIR principles used in data descriptions
Interoperable:
- FM-I1 Use a Knowledge Representation Language: https://doi.org/10.25504/FAIRsharing.jLpL6i
- FM-I2 Use FAIR Vocabularies: https://doi.org/10.25504/FAIRsharing.0A9kNV
Reusable:
- FM-R1.3 Meets Community Standards: https://doi.org/10.25504/FAIRsharing.cuyPH9
What are data descriptions?
Data descriptions document the meaning of each data attribute or variable in a dataset.
Depending on the discipline, similar documents may be called:
- codebooks
- data dictionaries
- labels or data tags
- data glossaries
Example:
| Variable name | Description | Scale |
|---|---|---|
| Weight | Weight of a human | kilograms |
| Height | Height of a human | centimetres |
| Age | Age of a human | years |
| Blood glucose level | Blood glucose level of a human | mg/dl |
Different fields use different names
No matter what the document is called, the goal is the same: describe what the dataset variables mean so others can interpret and reuse the data correctly.
How can data descriptions be reused?
Writing documentation takes time, so when possible you should reuse accepted community definitions instead of inventing a new description every time.
BioPortal is a good example of a registry where community-maintained ontology terms can be searched and reused:
- BioPortal: https://bioportal.bioontology.org/
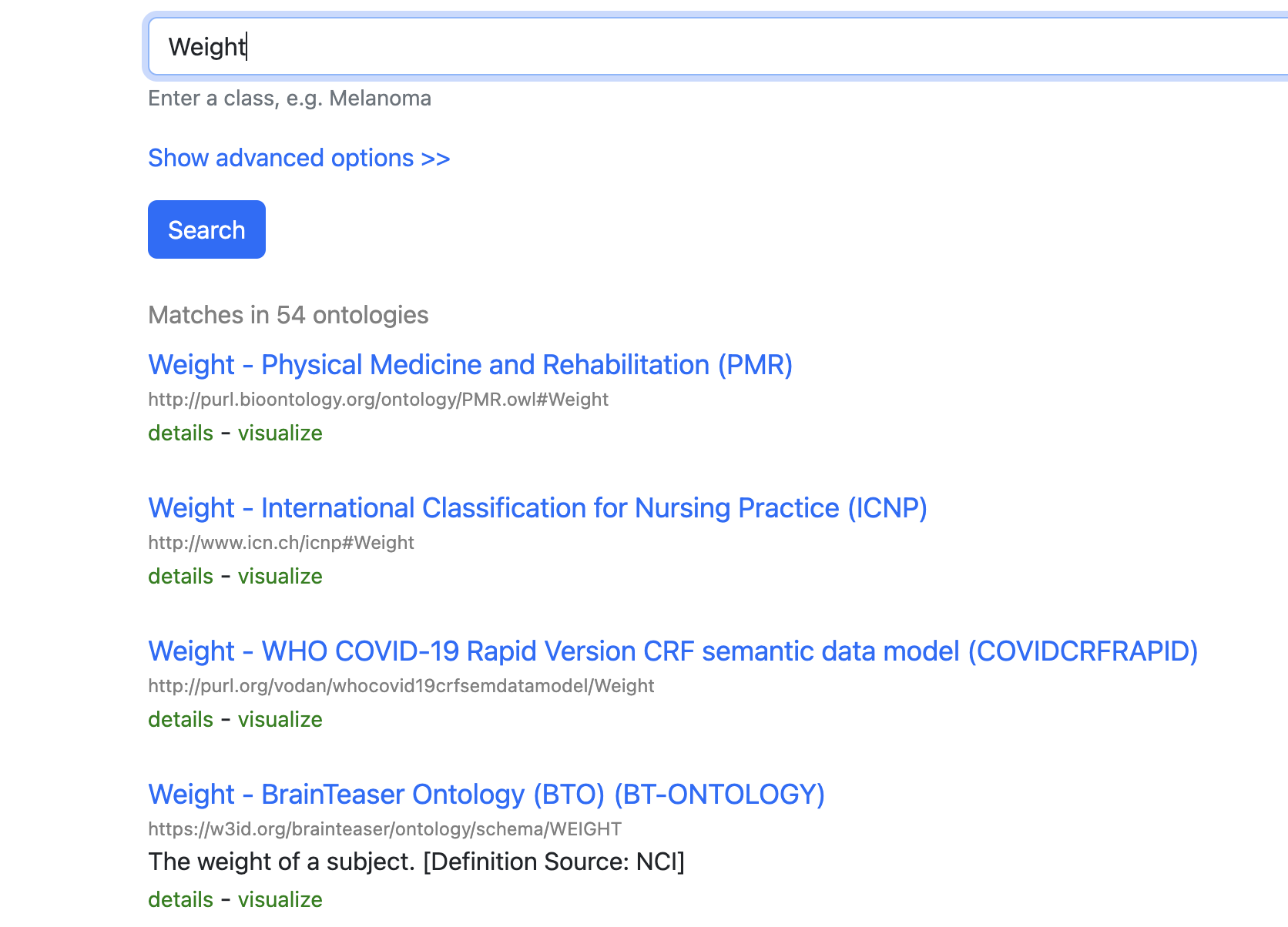

Reuse ontology terms when possible
Advantages:
- you do not need to redefine common concepts repeatedly
- you gain a persistent identifier for the concept
- other systems can align variables across datasets more easily
Disadvantages:
- sometimes no existing ontology fits the exact concept you need
Are there standard ways to create data descriptions?
There is no single universal format, but the minimum useful description usually includes:
- the variable name
- a definition
- ideally, a link to the reused ontology term or community definition
Some general-purpose vocabularies and metadata schemas include:
| Resource | Link | Use |
|---|---|---|
| Schema.org | https://schema.org/ | Generic web concepts |
| DBpedia | https://www.dbpedia.org/resources/lookup/ | Concepts derived from Wikipedia |
| DCAT | https://www.w3.org/TR/vocab-dcat-2/ | Data catalog concepts |
| Dublin Core | https://www.dublincore.org/specifications/dublin-core/dcmi-terms/ | General metadata terms |
Public registries that help locate vocabularies include:
- Linked Open Vocabularies: https://lov.linkeddata.es/dataset/lov/
- EU Vocabularies: https://op.europa.eu/en/web/eu-vocabularies
- BioPortal: https://bioportal.bioontology.org/
- AgroPortal: https://agroportal.lirmm.fr/
- EcoPortal: https://ecoportal.lifewatchitaly.eu/
- Ontology Lookup Service: https://www.ebi.ac.uk/ols/index
- Bioschemas: https://bioschemas.org/
Exercise
Visit BioPortal and search for a term describing
blood glucose level.
What ontology term would you reuse, and what persistent identifier does it provide?
There is more than one possible match, but a valid answer is to identify a term such as the clinical measurement concept and record its ontology URI or other persistent identifier provided by the registry.
Data descriptions are often maintained as tables in
.csv, .xlsx, or similar formats. For
databases, it is also useful to provide both a machine-readable schema
and a human-readable diagram.
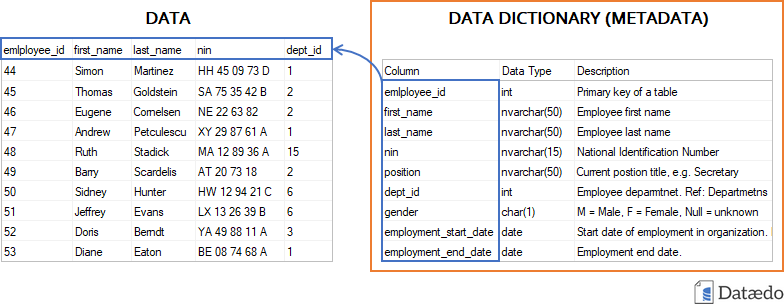
Other human-friendly approaches include dataset nutrition labels and automated codebook tools, but the FAIR principles push further toward structured, machine-actionable formats.
What is the relationship between data descriptions and linked data?
When data descriptions reuse ontology terms and stable identifiers, datasets can be transformed into linked data formats such as RDF and become easier to combine with other resources.
Tools that can help with this include:
| Tool | Source | GUI | Note |
|---|---|---|---|
| OpenRefine | https://openrefine.org/ | Yes | Flexible but can be heavy to install |
| RMLMapper | https://github.com/RMLio/rmlmapper-java/releases | No | Powerful, but technical |
| SDM-RDFizer | https://github.com/SDM-TIB/SDM-RDFizer | No | Requires programming familiarity |
| SPARQL-Generate | https://ci.mines-stetienne.fr/sparql-generate/ | Yes | Good if you want to learn SPARQL |
| Virtuoso Universal Server | https://virtuoso.openlinksw.com/ | Yes | Commercial licensing may apply |
| UM LDWizard | https://github.com/MaastrichtU-IDS/ldwizard-humanities | Yes | Quick route to publishable linked data |
Exercise
Transform a dataset from XLSX to RDF using UM LDWizard:
- Download the mock dataset: MOCK DATA
- Convert the dataset into RDF
- Record which ontology terms you reused for the variables
There is no single correct answer. The important part is to reuse existing ontology terms where possible and document the choices you made.
Scenario
Your marine biology group has discovered new organisms, but you cannot find an existing ontology that fits the concepts you need.
What should you do? Should you adapt an existing vocabulary, create local terms, or work with the community toward a new ontology extension?
- Data descriptions may appear under names such as codebook or data dictionary.
- Reusing ontology terms reduces ambiguity and improves interoperability.
- A useful description links dataset variables to accepted community concepts.
- Linked data becomes more feasible when descriptions are structured and identifier-based.
